Mycoplasma neophronis
(Suárez-Pérez et al., 2012)
Etymology
Gr. n. mukes – fungus, Gr. neut. n. plasma – anything formed, N.L. neut. n. Mycoplasma – fungus form; N.L. n. Neophron – a genus name of vultures, N.L. gen. n. neophronis – of Neophron
Taxonomy
Mycoplasmatales – Mycoplasmataceae – Mycoplasma – Mycoplasma neophronis (Hominis cluster), closely related to Mycoplasma spumans (16S rRNA gene sequence similarity – 98.70%) (Fig. 1)
Type strain
G.A.T (Canarian Egyptian vulture – Neophron percnopterus majorensis, Spain, 2003), (Fig. 2, 16S rRNA gene sequence)
Genomes
one draft genome (G.A.T – Spain) (NCBI Genome deposit per 11/05/2024)
Cell morphology
spherical – coccoid
Colony morphology
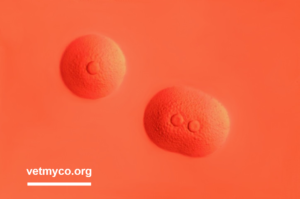
fried egg morphology (Fig. 3)
Metabolism
hydrolysis of arginine; non-fermentative, non-urea-hydrolyzing
Host
Canarian Egyptian vulture (Neophron percnopterus majorensis)
Habitat
upper respiratory tract
Disease(s)
unknown, no disease reported
Pathogenicity
factors unknown
Epidemiology
unknown, has once been isolated from 6 vultures in Gran Canaria, Spain
Diagnosis
cultivation and species identification by MALDI-ToF MS, serology or genetically
Fig. 1. Maximum likelihood tree showing the phylogenetic position of Mycoplasma neophronis G.A.T within the Hominis cluster of Mycoplasmataceae based on 16S rRNA gene sequences. The sequence of Mycoplasma synoviae WVU 1853T was used as out-group (Synoviae cluster). Numbers at nodes represent bootstrap confidence values (1000 replications). Only values > 80% are shown. Bar, number of substitutions per nucleotide position. Credits: Joachim Spergser (Vetmeduni Vienna)
CTGGCTGTGTGCCTAATACATGCATGTCCGAGCGAGGTTCTTCGGAACCTAGCGGCGAATGGGTGAGTAACACGTGCTTAATCTACCCTTCAGATTGGAATACCCAATGGAAACATTGGCTAATGCCGGATACGCATGGAATCGCATGATTCCGTTGTGAAAGGGGCCTTTAAAGCCCCACTGAAGGATGAGGGTGCGGAACATTAGTTAGTTGGTAGGGTAATGGCCTACCAAGACTATGATGTTTAGCCGGGTCGAGAGACTGAACGGCCACATTGGGACTGAGATACGGCCCAAACTCCTACGGGAGGCAGCAGTAGGGAATATTCCACAATGAGCGAAAGCTTGATGGAGCGACACAGCGTGCACGATGAAGGTCTTCGGATTGTAAAGTGCTGTTATAAGGGAAGAACACTACATTGAGGAAATGCTTTGTAGCTGACGGTACCTTGTCAGAAAGCGATGGCTAACTATGTGCCAGCAGCCGCGGTAATACATAGGTCGCAAGCGTTATCCGGAATTATTGGGCGTAAAGCGTTCGTAGGCTGTTTATTAAGTCTGGAGTCAAATCCCGGGGCTCAACCCCGGCTCGCTTTGGATACTGATAAACTAGAGTTGGATAGAGGTAAGCGGAATTCCATGTGAAGCGGTGAAATGCGTAGATATATGGAAGAACACCAAAGGCGAAGGCAGCTTACTGGGTCTATACTGACGCTGAGGGACGAAAGCGTGGGGAGCAAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACGATGATCATTAGTCGGTGGAGAGTTCACTGACGCAGCTAACGCATTAAATGATCCGCCTGAGTAGTATGCTCGCAAGAGTGAAACTTAAAGGAATTGACGGGGACCCGCACAAGCGGTGGAGCATGTGGTTTAATTTGAAGATACACGGAGAACCTTACCCACTCTTGACATCTTCCGCAAAGCTATAGAGATATAGTGGAGGCTAACGGAATGACAGATGGTGCATGGTTGTCGTCAGCTCGTGTCGTGAGATGTTTGGTCAAGTCCTGCAACGAGCGCAACCCCTATCTTTAGTTACTAACGAGTCATGTCGAGGACTCTAGAGATACTGCCTGGGTAACCGGGAGGAAGGTGGGGATGACGTCAAATCATCATGCCTCTTACGAGTGGGGCCACACACGTGCTACAATGGTCGGTACAAAGAGAAGCAATATGGCGACATGGAGCAAATCTCAAAAAGCCGATCTCAGTTCGGATTGGAGTCTGCAATTCGACTCCATGAAGTCGGAATCGCTAGTAATCGCAGATCAGCTATGCTGCGGTGAATACGTTCTCGGGTCTTGTACACACCGCCCGTCACACCATGGGAGCTGGTAATACCCAAAGTCGGTTTGCTAACCTCGGAGGCGACCGCCTAAGGTAGGACTGGTGACTGGGGTGAAGTCGTAACAAGGT
Fig. 2. 16S rRNA gene sequence of Mycoplasma neophronis G.A.T (Accession number: NR_108494)Fig. 3. Colonies of Mycoplasma neophronis G.A.T on modified Hayflick’s agar after 4 days of incubation exhibiting fried egg morphology. Note, colour change of solid medium from ochre to reddish based on release of ammonia resulting from hydrolysis of arginine creating an alkaline pH. Bar, 1 mm. Credits: Joachim Spergser (Vetmeduni Vienna)